 Conserved regulatory strategies involving combinations of transcription factors (TFs) establish unique neuronal identities ( Allan and Thor, 2015 Enriquez et al., 2015 Hobert, 2016). Indeed, neuronal subtypes express highly diverse repertoires of CSPs during circuit assembly ( Tan et al., 2015 Li et al., 2017 Sarin et al., 2018). Gain and loss of function genetic studies have shown that combinations of different CSP families regulate this specificity ( Zarin et al., 2014). Studies in both vertebrates and invertebrates have led to the identification of cell surface proteins (CSPs) that mediate selective association between neurites ( Tessier-Lavigne and Goodman, 1996 de Wit and Ghosh, 2016 Zinn and Özkan, 2017). 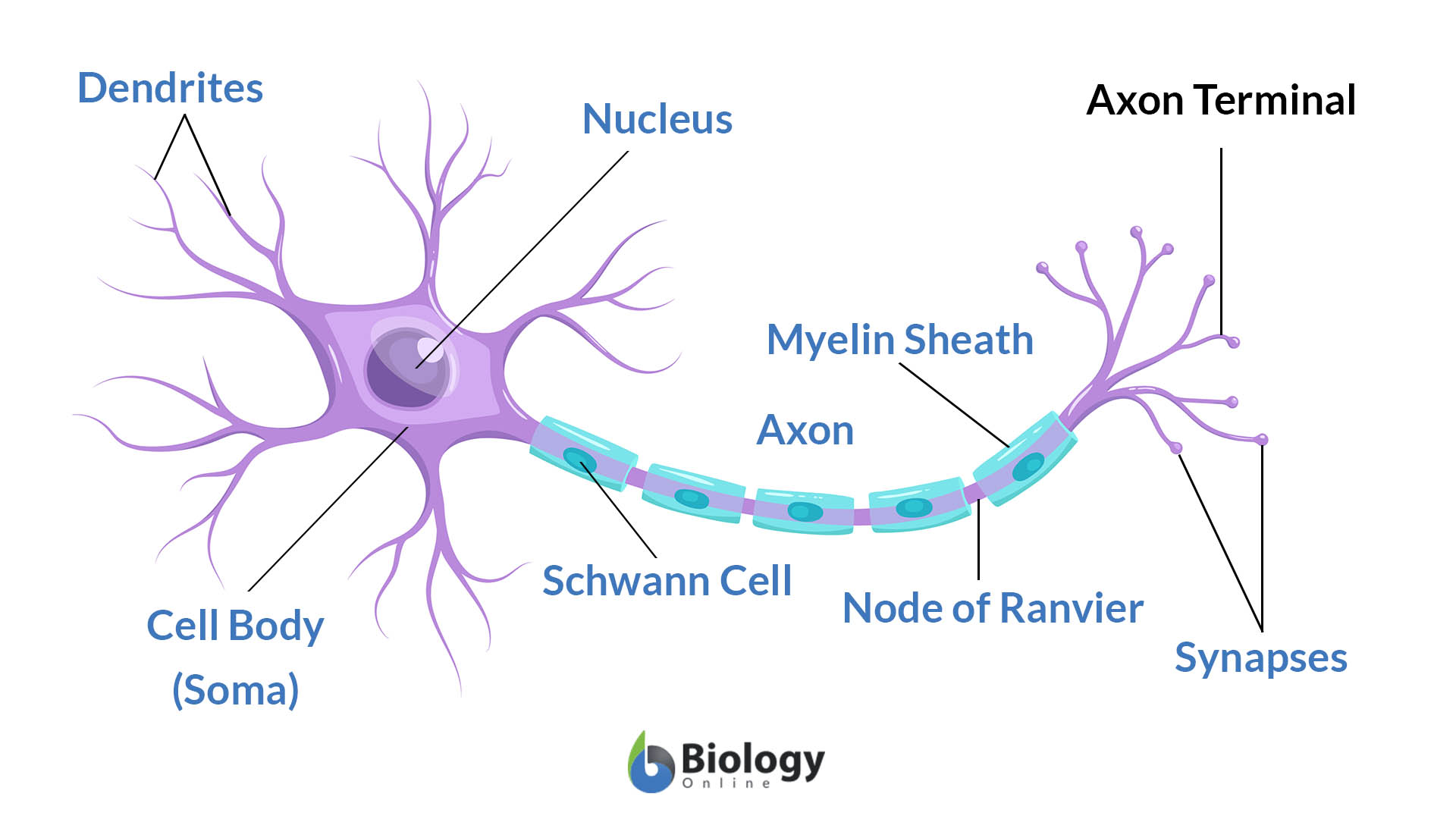
Vast numbers of neurites from a diversity of neurons are intermingled within the developing central nervous system, and they form highly specific synaptic connections with a discrete subset of the neurons they contact. Much of the specificity of inputs and outputs of neurons in the mammalian CNS is also genetically determined ( Sanes and Zipursky, 2010). In invertebrates, stereotypical wiring patterns are genetically encoded in the programs regulating the development of neurons. At the cellular level, this entails each neuron adopting a specific wiring pattern, the combination of specific synaptic inputs and outputs. We propose that modular transcriptional programs for distinct wiring features are assembled in different combinations to generate diverse patterns of neuronal connectivity.īrain function relies on precise patterns of synaptic connections between neurons. Gain and loss of function studies provide evidence for independent control of different wiring features. These programs were defined by the expression of a few transcription factors and different combinations of cell surface proteins. Single-cell profiling during development revealed distinct transcriptional programs defining each dendrite and axon wiring pattern. 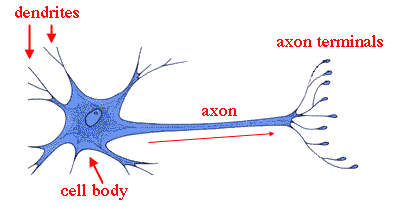
Eight subtypes of T4/T5 neurons are defined by combinations of two patterns of dendritic inputs and four patterns of axonal outputs. Here, we focused on Drosophila T4/T5 neurons, a class of closely related neuronal subtypes with different wiring patterns. Nevertheless, a clear logic underlying the transcriptional control of neuronal connectivity has yet to emerge. Single-cell RNA sequencing has revealed a vast transcriptional diversity of neurons. Patterns of synaptic connectivity are remarkably precise and complex. 
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
